- Nepal’s Centre for Molecular Dynamics has developed a DNA reference database containing genetic and geographic information on 120 of the country’s estimated 200 wild tigers.
- Law enforcement officials used the database to identify the species, sex, and estimated geographic origin of confiscated animal parts suspected to be tigers, pinpointing most of them to individual national parks.
- Such databases have the potential to support not only forensics, but also disease research and monitoring population dynamics, particularly if countries can share genetic data.
In December 2015, officials from Nepal’s Central Investigation Bureau had a hunch that something was amiss with the country’s tigers. The bureau, in the past few months, had seized more than 15 suspected tiger parts (13 pieces of skin and two blood-smeared knives) from various areas of Nepal.
Their main challenge was to identify these pieces of evidence with confidence, to know whether they actually came from tigers or not. But more importantly, they wanted leads to tap into wildlife criminal networks active in the country. Observing the stripes on the skin could only provide limited clues. That was when they decided to test a new ‘tell-tale’ technology developed by a research organisation based in the capital Kathmandu.
“They knew of our tiger DNA database, and they wanted to seek our help in identifying the species of seized parts and possibly track the origin of these putative tiger parts,” said Dr Dibesh Karmacharya, director at Centre for Molecular Dynamics-Nepal(CMDN).
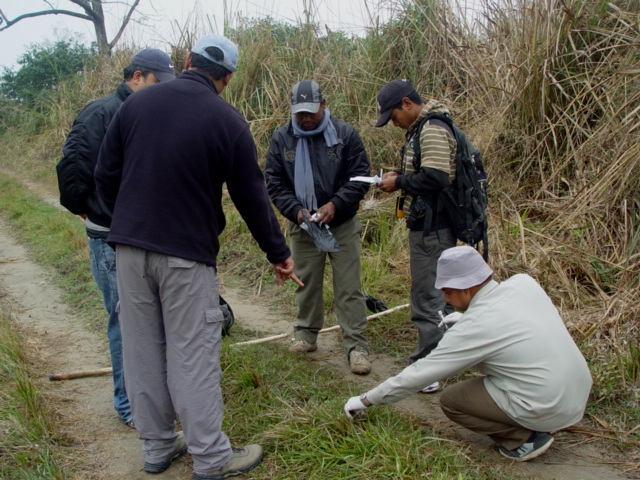
Work on the database, which took two years to develop, started in 2011 when field biologists and conservation geneticists roamed the woods of Nepal’s southern plains (the Terai Arc Landscape) to collect tiger DNA. “As we walked along trails in the jungle, we stopped every few minutes and stooped down to scoop any possible tiger faecal sample—they look like beads,” said Prajwol Manandhar, CMDN’s bioinformatics specialist.
The work was done as part of CDMN’s USAID-funded Nepal Tiger Genome Project, Nepal’s first molecular-based wildlife research program that would establish region’s most comprehensive DNA database of one of the world’s most endangered cats.
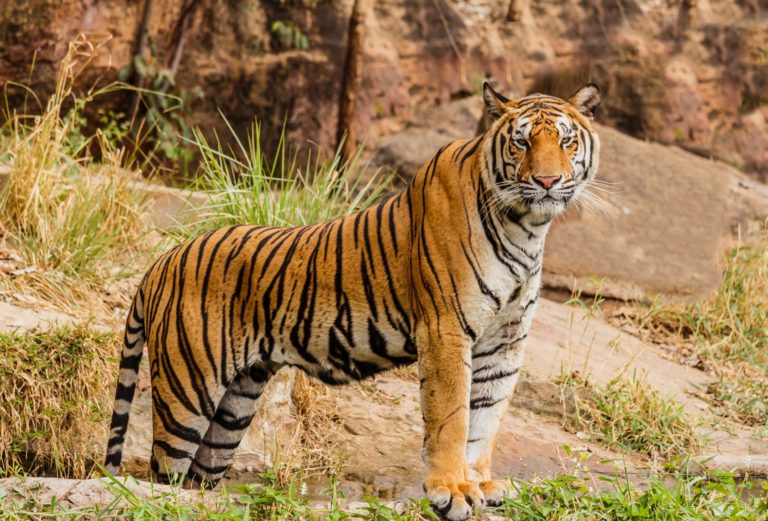
The number of Bengal tigers, which in Nepal are found in fragmented habitats in the country’s southern plains (the Terai Arc Landscape), has been dwindling–mainly due to poaching. Nepal is also a known wildlife crime transit between India and China for traders in tiger parts (skin, bones, teeth and reproductive organs, among others) used as traditional medicine and/or commodities in Southeast Asia, China, US and Europe.
Until now, authorities relied on visual methods to keep track of tigers in Nepal’s protected areas, with limited success. But all that is changing.
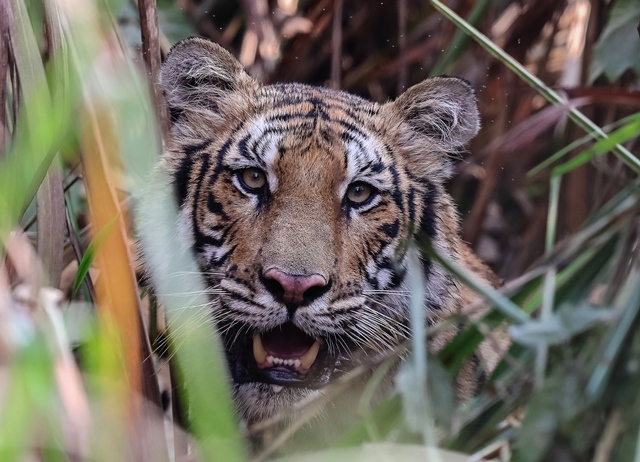

“The novel idea of identifying and tracking wildlife parts is based on DNA,” Karmacharya said. “However, the key component of the whole process relies heavily on a complete reference DNA database. When faeces pass through the intestines of an animal, it contains worn-out cells that carry the animal’s genetic blueprint (the DNA). So when we are collecting faeces, we are actually collecting sources of DNA of that particular animal.”
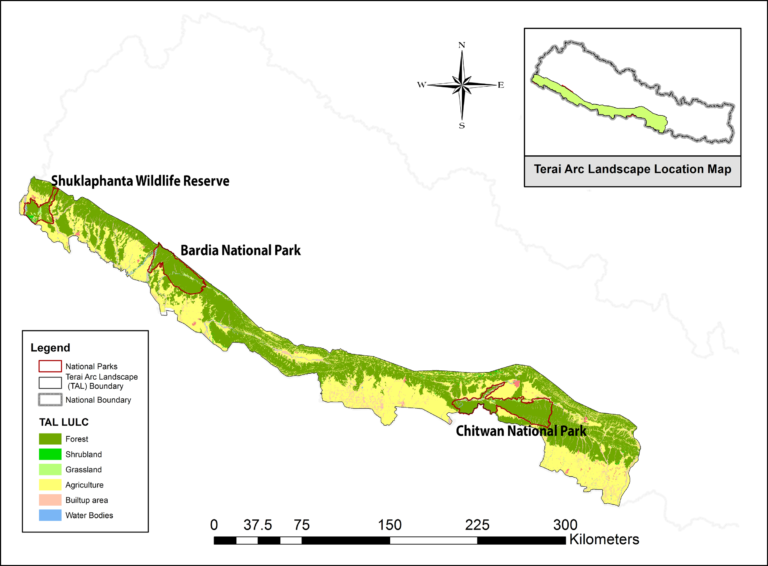
“We analyzed 770 different fecal samples from Chitwan, Bardia [a.k.a. Bardiya], Shuklaphanta national parks and Parsa Wildlife Reserve, along with known wildlife corridors, to build individual DNA fingerprint profiles of 120 out of a total estimated 200 tigers living in Nepal,” said Adarsh Man Sherchan, one of the scientists involved in the project.
The profiles contain each animal’s nuclear DNA microsatellite genotype and sex data, which they analyzed to identify the species, sex and likely geographic location of the sample’s origin.

When the call from the Nepal Police came, scientists at CDMN were excited to test the applications of the database they had built.
The results were unmistakable. “We found out that all 15 samples were indeed from tigers–10 of the specimens were from male tigers and five from female,” Manandhar said. The group recently published their findings in the scientific journal PLOS One.
Their method allowed the scientists to find an exact match with individuals in the database or pinpoint the source population of the poached parts. “So far, only Nepal has this capability,” Karmacharya said.
“One of the seized tiger samples assigned to Bardia was a 100 percent match with a female tiger’s DNA in our database. This shows that the tigress roamed her territory while we were sampling. And she was killed sometime between 2013-2015,” said Jyoti Joshi, Wildlife Programme Manager at CMDN.
Although Joshi and her teammates were saddened by the news, they were also gratified that the incident proved that the method worked.
Meanwhile, DNA fingerprints of the samples from the tiger parts received from the police were subjected to a host of algorithms to trace the origin of the samples. “By utilizing four matching tools, six samples were assigned to Bardia National Park. Five samples could not be assigned to Nepal’s tiger populations,” Sherchan said.
The unusually high number of poached tiger parts from Bardia alarmed law enforcement officials, and there was a greater realization that the national park in mid-west Nepal was becoming a hub for tiger poachers.
“After we found this, we deployed special teams to track down the poachers and made some arrests,” said Pravin Pokherel, an official who was working for CIB at the time of the investigation.
Nepali officials identified a nomadic group from Rajasthan, India as one of the active poaching networks, monitored their activities and subsequently stopped them.
The officials hope that similar action can be taken in the future when tiger parts are recovered from poachers. They also hope that following the developments, laws are amended to allow DNA evidence to be presented in courts in criminal cases related to poaching.
The researchers believe that the five samples that could not be assigned to Nepal’s tiger population most likely came from nearby tiger range countries (Bhutan, India or Bangladesh). Tigers commonly move among these countries.
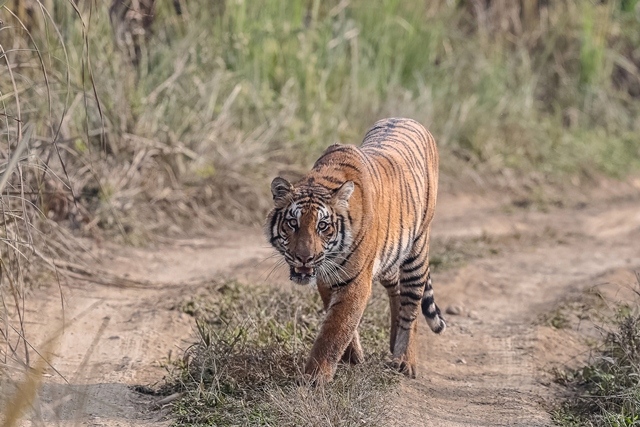

“However, due to a lack of transboundary genetic data sharing between the neigbouring countries, the challenge of identifying or linking non-native seizures to its source population still remains largely ineffective,” Karmacharya said. “The need for transboundary genetic data sharing is critical and almost imperative if forensic analysis is to become regionally functional and practically viable to save this precious animal.”
The Tiger Genome Project and the DNA reference database it created have the potential to support not only forensics, but also disease research and monitoring population dynamics, among other unexplored applications.
“This is an exciting time for tiger researchers in Nepal,” Sherchan said. “However, we need to tread with caution. The data collected has a shelf life, and further rounds of studies are necessary to ensure that we have an up-to-date database to save the Bengal tiger.”
This article was originally published on Mongabay. Read the original article.